In a previous article, I began a series of four responses to YouTuber Dave Farina (aka “Professor Dave”) about his video reviewing Dr. Michael Behe’s three books. Here I will turn my attention to Mr. Farina’s comments regarding bacterial flagella.
In relation to the flagellum, the video complains about Behe’s “dishonest usage of terminology pertaining to machinery,” including phrases such as “outboard motor,” “drive shaft,” “universal joint,” “bushings,” and “clutch and braking system.” In reality, this terminology is used widely in the scientific literature. It’s not unique to Behe. On the contrary, in reference to flagella, the literature is full of such terms including “motor”, [1] “drive shaft,” [2] “universal joint,” [3] “bushing,” [4] and “clutch.” [5] The word “machine” itself has a wide circulation. [6] Is Farina going to charge the entire flagella research community with dishonesty as well?
Co-option Scenarios for the Origins of Bacterial Flagella
According to the video, “A flagellum that merely twitches instead of rotating smoothly would also produce motion and thus could be selected for.” But a type IV pilis, which enables twitching motility (a form of bacterial translocation over moist surfaces), is very different from a flagellum. Twitching motility occurs by extension, tethering, and retraction of the type IV pilus, which functions in a manner akin to a grappling hook. A flagellum, on the other hand, rotates as it is driven by a proton motive force across the cell membrane. The assembly mechanisms of pili and flagella are also quite different.
The video complains that Behe fails to acknowledge the existence of alternative flagellar systems that are simpler than the model system found in Salmonella species and Escherichia coli. However, the fact that an alternative system lacks a specific component that is essential in another system does not mean that the former lacks an alternative mechanism for achieving the same outcome. The most robust concept of irreducible complexity understands it as a property of a system that is contributed to by multiple subfunctions, the removal of one of which causes the overall system to effectively cease performing its job. Note that each individual subfunction could, in principle, be performed by multiple protein components. Likewise, a single protein component could perform more than one of those subfunctions. Furthermore, the identity of the specific components performing each respective subfunction could differ from one organism to the next. It is therefore not the identity of the structural parts that is important in an irreducibly complex system, but rather the essential functions that need to be performed in order for a higher-level objective to be realized.
Moreover, pointing to homologues of flagellar proteins does not undermine the argument from irreducible complexity, since co-opting those proteins to produce a flagellar system requires multiple co-incident changes in order for the new system to be realized. For example, flagellar-specific proteins would not confer a selective advantage until incorporated into the flagellar system. But the necessary proteins that serve roles in other systems will not become incorporated into the flagellar system before these flagellar-specific proteins arise. This is quite aside from the need to have complementary protein-protein binding interfaces, as well as a choreographed assembly system to ensure that the proteins are assembled in the appropriate order.
Resurrecting a Flagellum
In a 2016 article at Evolution News, Behe asks,
[W]hy doesn’t [Kenneth Miller] just take an appropriate bacterial species, knock out the genes for its flagellum, place the bacterium under selective pressure (for mobility, say), and experimentally produce a flagellum — or any equally complex system — in the laboratory? (A flagellum, after all, has only 30-40 genes, not the hundreds Miller claims would be easy for natural selection to rapidly redesign.) If he did that, my claims would be utterly falsified. But he won’t even try it because he is grossly exaggerating the prospects of success.
The video by Farina comments,
Hilariously, [Behe] is oblivious to the fact that this precise experiment was carried out the year before. Here’s the paper. Gene deletion produced two strains of bacteria with no flagellum. They then introduced selective pressure for motility by depleting the nutrients in the colony. Within 96 hours, both strains had regenerated flagellar motility by a pathway involving two successive point mutations in genes that served other purposes.
However, the paper that Farina cites7 does not do this at all. Not for the first time with this video, I wonder if he has in fact read the paper. All that the researchers deleted was the flagellar master switch protein, FleQ, in Pseudomonas fluorescens. After a few days of incubating the bacterial cells on Petri dishes, they reacquired their ability to grow flagella. The genetic basis for this reactivation of the flagella is that another master switch protein, NtrC, that is a structurally similar homolog of FleQ — responsible for turning on genes involved in nitrogen metabolism — already had the ability, to some extent, to cross-bind to the promoter usually bound by FleQ. When produced in excess, as a result of a broken regulator, NtrC was thus able to drive flagellar synthesis. As a consequence of this mutation, the bacterial cell lost its ability to regulate its nitrogen metabolism genes. An article in The Scientist describes this research:
But while the re-evolved flagella enabled the bacteria to access food supplies at the farthest reaches of the Petri dish, the ability came at a price. ‘The bacteria that became much better at swimming were much worse at nitrogen regulation,’ said Johnson. However, she added, ‘sometimes the advantage can be so great that it’s worth paying that cost because otherwise you die.’
Thus, contrary to the Farina video’s claims, this paper does not document the de novo evolutionary origins of a bacterial flagellum at all — far from it. In fact, Behe has already addressed the paper here.
The Waiting Times Problem
In 2004, Michael Behe and David Snoke published a paper in the journal, Protein Science. [8] About this paper, Farina has three complaints. The first complaint is that, “Behe and Snoke found that the target sequence did actually evolve, in population sizes and timeframes that are entirely realistic, and if anything, quite small compared to real-world populations. The paper literally proves them wrong and they somehow count it as a win anyway.” Farina mentions Behe’s expert testimony at the 2005 Kitzmiller v. Dover trial:
When questioned about his 2004 paper, Behe tacitly acknowledged that the population size in their model was orders of magnitude smaller than real-world bacterial populations, which had the effect of vastly underestimating the rate at which such “irreducible” traits could evolve… In one striking exchange, Behe acknowledged a paper which indicated that there are more prokaryotes in a single ton of soil than in his model population, and that there is a lot more than one ton of soil on Earth.
However, this objection stems from Farina’s misreading of the paper. As Behe himself explains in the very transcript that Farina cites, “forming a new disulfide bond might require as few as two point mutations. But forming other multi-residue features such as protein-protein binding sites might require more.” The graph below (figure 6 of the paper) shows Behe and Snoke’s estimate of the time to fixation (along the y-axis) versus the number of substitutions needed for a new feature to evolve (along the x-axis). On the top axis, values for the needed population sizes are given. The point is that, as the number of needed co-dependent mutations increases, so too does the needed population size and waiting time to fixation.
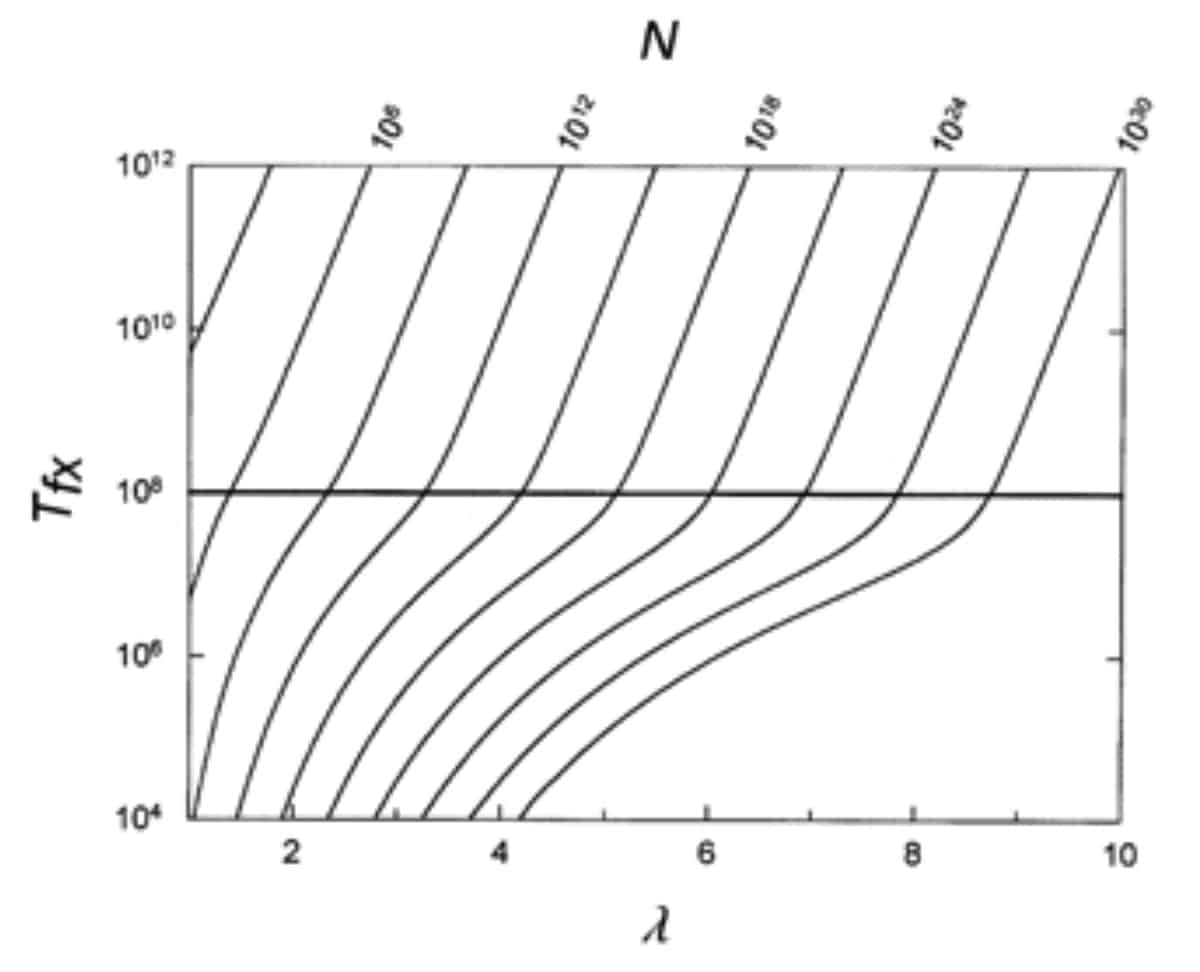
As Behe and Snoke explain in the paper, in a scenario where three substitutions are required for a novel feature to arise, a population size of roughly 1011 individuals is necessary for it to become fixed over the course of 108 (108 generations is marked as a horizontal bar on the figure). If the complex trait in question requires even more substitutions, it would require considerably more time. If six mutations were needed, the average population size required for it to become fixed in 108 generations would be on the order of 1022 individuals. Given that 1030 is a plausible estimate of the number of microorganisms on the entire planet [9], these numbers become prohibitive very quickly.
The second complaint is that, “In their model, Behe and Snoke permitted only single-base mutations and natural selection — no recombination, no duplications beyond the initial presumed one, no other evolutionary changes.” But the authors explicitly say that “Because the model presented here does not include recombination, the results can be considered to be most applicable to a haploid, asexual population.” Nonetheless, they do note in the conclusion to their paper that “the results also impinge on the evolution of diploid sexual organisms,” since large multicellular organisms have much, much smaller population sizes than bacteria. If the evolution of complex features is difficult for microorganisms (with their massive population sizes and short generation turnover times), how much more so for large animals? Though one might counter, in the case of diploid sexual species, that recombination allows for neutral mutations to occur separately in a population and to later combine by sexual recombination, Christiansen et al. have shown, in a paper published in Theoretical Population Biology, that “Recombination lowers the waiting time until a new genotypic combination first appears, but the effect is small compared to that of the mutation rate and population size” (emphasis added). [10]
Finally, Farina complains that “They also specified a pre-determined target sequence and only considered the simulation to have been ‘successful’ if that specific target evolved.” But this is incorrect. Rather, the paper provides estimates for how many organisms would be required, and over how long a time frame, for multiple co-dependent mutations (none of which by themselves confers an advantage) to become fixed in a population.
Next, I will respond to Farina’s criticisms of Dr. Behe’s The Edge of Evolution.
Footnotes
1. Minamino T, Imada K, Namba K. Molecular motors of the bacterial flagella. Curr Opin Struct Biol. 2008; 18(6):693-701.
2. Johnson S, Furlong EJ, Deme JC, Nord AL, Caesar JJE, Chevance FFV, Berry RM, Hughes KT, Lea SM. Molecular structure of the intact bacterial flagellar basal body. Nat Microbiol. 2021; 6(6):712-721.
3. Kitao A, Hata H. Molecular dynamics simulation of bacterial flagella. Biophys Rev. 2018; 10(2):617-629.
4. Yamaguchi T, Makino F, Miyata T, Minamino T, Kato T, Namba K. Structure of the molecular bushing of the bacterial flagellar motor. Nat Commun. 2021 Jul 22;12(1):4469.
5. Blair KM, Turner L, Winkelman JT, Berg HC, Kearns DB. A molecular clutch disables flagella in the Bacillus subtilis biofilm. Science. 2008;320(5883):1636-8.
6. Sowa Y, Berry RM. Bacterial flagellar motor. Q Rev Biophys. 2008 May;41(2):103-32.
7. Taylor TB, Mulley G, Dills AH, Alsohim AS, McGuffin LJ, Studholme DJ, Silby MW, Brockhurst MA, Johnson LJ, Jackson RW. Evolution. Evolutionary resurrection of flagellar motility via rewiring of the nitrogen regulation system. Science. 2015; 347(6225):1014-7.
8. Behe MJ, Snoke DW. Simulating evolution by gene duplication of protein features that require multiple amino acid residues. Protein Sci. 2004; 13(10):2651-64.
9. Whitman WB, Coleman DC, Wiebe WJ. Prokaryotes: the unseen majority. Proc Natl Acad Sci U S A. 1998; 95(12):6578-83.
10. Christiansen FB, Otto SP, Bergman A, Feldman MW. Waiting with and without recombination: the time to production of a double mutant. Theor Popul Biol.1998;53(3):199-215.
Note: This essay has been adapted from a blog post originally published at Evolution News & Science Today on June 26, 2023.

2 thoughts on “Answering Farina on Behe’s Work: Bacterial Flagella”
Pingback: Answering Farina on Behe’s Work: The Edge of Evolution - Jonathan McLatchie | Writer, Speaker, Scholar
Pingback: Answering Farina on Behe’s Work: Darwin Devolves - Jonathan McLatchie | Writer, Speaker, Scholar
Comments are closed.